Host control over ineffective (non fixing) rhizobial symbionts
Host control
mechanisms are thought to be critical for selecting against ineffective mutants in
symbiont populations. In rhizobia, non-fixing strains are common and these ineffective symbionts are predicted to destabilize the interaction. Some of our recent research has tested a legume host’s
ability to constrain the infection and proliferation of a native ineffective
rhizobial strains.  Lotus strigosus hosts were experimentally inoculated with pairs of Bradyrhizobium strains that naturally
vary in symbiotic benefit, including a cheater symbiont strain that
proliferates in the roots of singly-infected hosts yet provides zero growth
benefits. Within coinfected hosts, the cheaters (and other ineffective strains) exhibited lower infection rates
than competing beneficial strains and grew to smaller population sizes within
those nodules. In vitro assays
revealed that infection-rate differences among competing strains were not due
to variation in rhizobial growth rate or inter-strain toxicity. These results
can explain how a rapidly growing cheater symbiont – that exhibits a massive
fitness advantage in single infections – can be prevented from sweeping through
a beneficial population of symbionts.
Lotus strigosus hosts were experimentally inoculated with pairs of Bradyrhizobium strains that naturally
vary in symbiotic benefit, including a cheater symbiont strain that
proliferates in the roots of singly-infected hosts yet provides zero growth
benefits. Within coinfected hosts, the cheaters (and other ineffective strains) exhibited lower infection rates
than competing beneficial strains and grew to smaller population sizes within
those nodules. In vitro assays
revealed that infection-rate differences among competing strains were not due
to variation in rhizobial growth rate or inter-strain toxicity. These results
can explain how a rapidly growing cheater symbiont – that exhibits a massive
fitness advantage in single infections – can be prevented from sweeping through
a beneficial population of symbionts.
Regus, J.
U., K.A. Gano, Holllowell, A.C., V. Sofish, J. L.
Sachs . Lotus hosts delimit the
mutualism-parasitism continuum of Bradyrhizobium
2015 Journal of Evolutionary
Biology. 28, 447-458
[PDF]
Regus, J. U., Gano, K. A., Hollowell, A. C., Sachs, J. L. Efficiency of partner choice and sanctions in Lotus is not altered by nitrogen fertilization 2014 Proceedings of the Royal Society of London. 281, 20132587 [PDF]
Sachs, J.L., Russell, J. E., Lii, Y. E.,
Black, K. C., Lopez, G., and Patil, A. S. 2010. Host control over infection and
proliferation of a cheater symbiont. Journal of Evolutionary Biology.
23:1919-1927. [PDF]
Simms, E. L., Taylor, D. L., Povich, J., Shefferson, R. P., Sachs, J. L., Urbina, M., and Tauszick, Y. 2006. An empirical test of partner choice mechanisms in a wild legume- rhizobium interaction. Proceedings of the Royal Society of London 273:77-81.[PDF]
The evolution
of uncooperative symbionts
A
key prediction for symbioses is that they are evolutionarily
unstable: mutants are predicted to arise and spread in symbiont populations
that exploit host resources without paying costs to hosts. For instance, rhizobia
are bacteria that fix nitrogen in legume roots in exchange for photosynthates
from their hosts. Uncooperative rhizobia – including non-fixing and
non-nodulating strains – appear common in agriculture, yet their population
biology and origins remain unknown in natural soils. In a recent study (Sach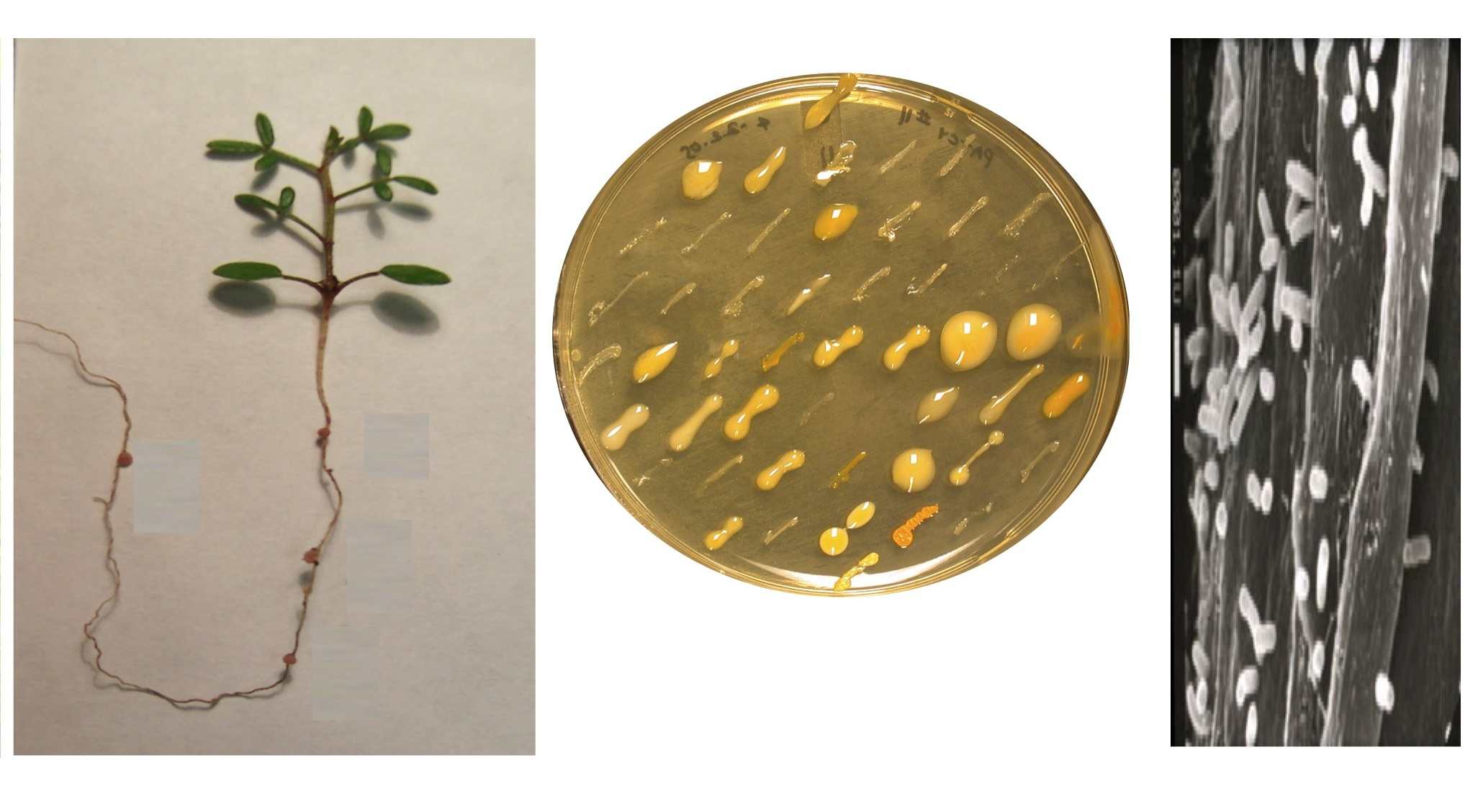 s et
al. 2010a, below), a phylogenetically broad sample of 62 wild-collected
rhizobial isolates was experimentally inoculated onto Lotus strigosus to assess
their nodulation ability and effects on host growth.
s et
al. 2010a, below), a phylogenetically broad sample of 62 wild-collected
rhizobial isolates was experimentally inoculated onto Lotus strigosus to assess
their nodulation ability and effects on host growth.
A cheater strain was
discovered that proliferated in host tissue
while offering no benefit; its fitness was superior to that of beneficial
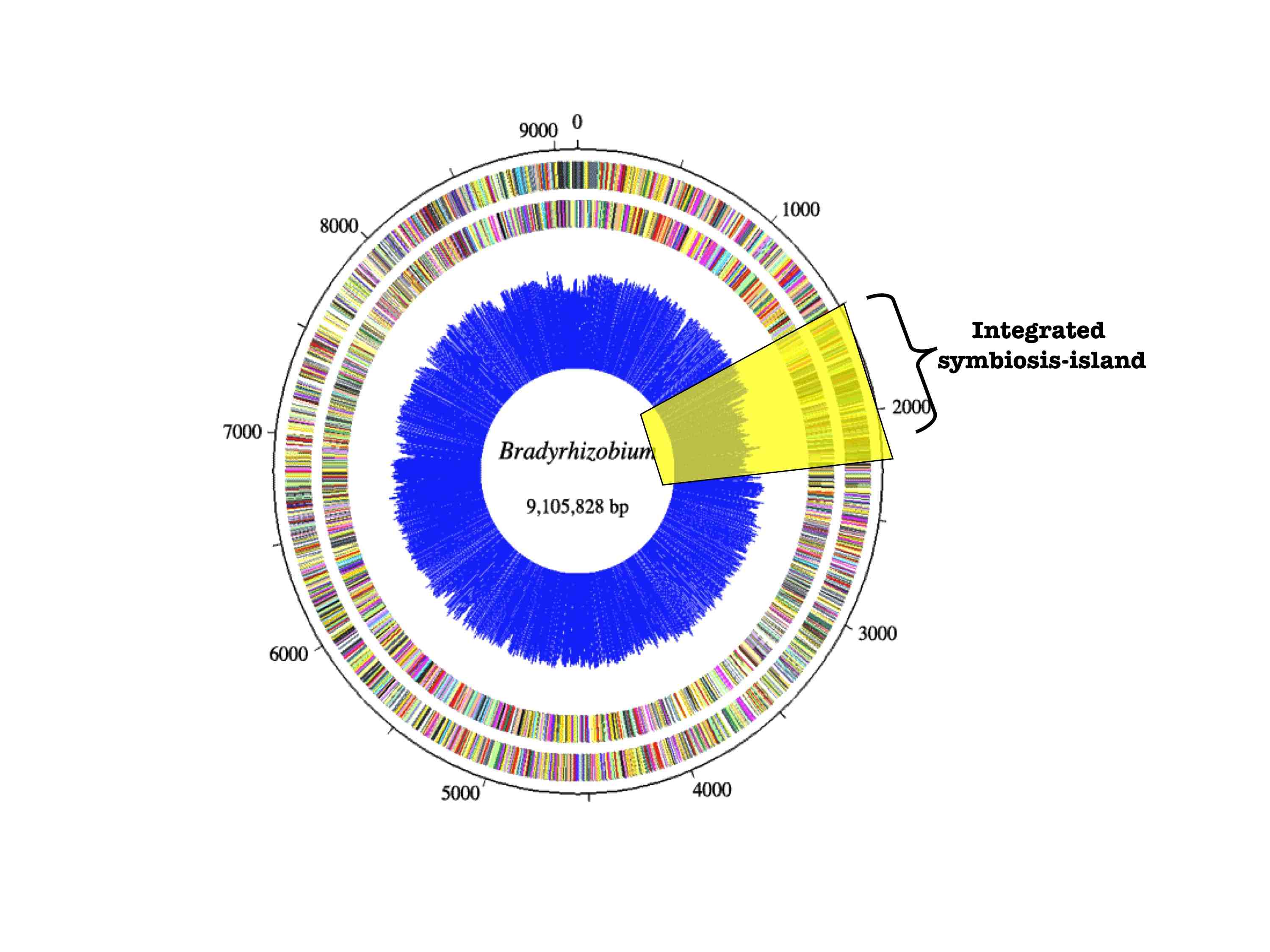
strains. Phylogenetic reconstruction of Bradyrhizobium
rDNA and transmissible symbiosis-island loci suggest that the cheater evolved
via a massive symbiotic gene transfer event. Many non-nodulating strains were
also identified and it appears that nodulation ability has been recurrently
lost in the symbiont population.
Sachs,
J.L., Ehinger, E. O., & Simms, E. L. 2010a. Origins of cheating and loss of symbiosis in wild Bradyrhizobium. Journal of Evolutionary Biology. 23:1075-1089. [PDF]
Sachs, J. L., and Simms, E.L. 2008. The origins of uncooperative
rhizobia. Oikos117:961-966. [PDF]
Sachs, J. L., and Simms, E.L. 2006.
Pathways to mutualism breakdown. Trends
in Ecology and Evolution 21:585-592 [PDF]
Sachs, J. L. and Wilcox, T.P. 2006. A
shift to parasitism in the jellyfish symbiont Symbiodinium microadtriaticum. Proceedings of the Royal Society of
London, B. 273:425-429. [PDF]
Bacterial symbioses range from short-term interactions to
bacterial-derived organelles and endosymbionts. Despite the
importance and near ubiquity of these beneficial infections, key questions
about evolution of bacterial symbioses remain unanswered. One fundamental problem is to resolve the
origins of symbiotic traits in bacteria. Originally, theoreticians predicted that pathogenic bacteria often undergo transitions into commensal or
beneficial partners. More recently biologists have used genomic data to argue
that parasites and symbionts must represent independent origins (of host
association) because genetic constraints hinder transitions between these
states. A related problem is to test hypotheses about the evolutionary robustness
of bacterial symbioses. Cooperation models predict that faster-evolving microbes are selected to exploit eukaryotic hosts (hence that parasitic mutants can fix in symbiont
populations). Nonetheless studies of bacterial lineages have rarely studied
transition rates between symbiosis and parasitism or examined the stability of
bacterial cooperation over deep time.
Sachs, J. L. and Bull, J.J. 2005.
Experimental evolution of conflict mediation between genomes Proceedings of the National Academy of
Sciences 102:390-395. [PDF]
Medina, M. and Sachs, J.L. 2010. Symbiont genomics; Our new tangled bank. Genomics 95:129-137. [PDF]
Sachs, J. L., Essenberg, C. and Turcotte. M. M. 2011 New Paradigms for the evolution of beneficial infections. Trends in Ecology and Evolution. 26: 202-209. [PDF]
Sachs, J. L., Russell, J. E. and Hollowell, A. C. 2011. Evolutionary instability of symbiotic
function in Bradyrhizobium. PLoS One 6: e26370. [PDF]
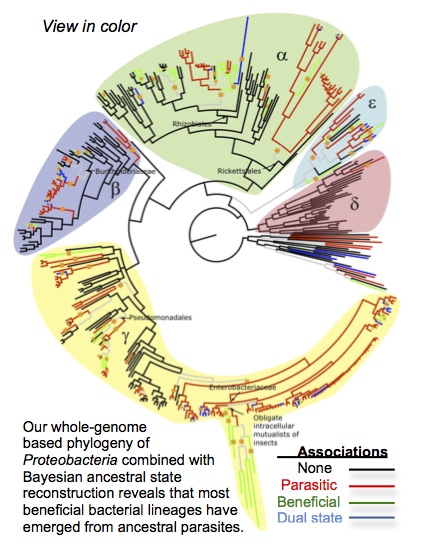 Sachs, J.L., Skophammer, R.G., and Regus, J.U. 2011. Evolutionary transitions in bacterial symbiosis. Proceedings of the National Academy of Sciences USA. 108: 10800-10807. [PDF]
Sachs, J.L., Skophammer, R.G., and Regus, J.U. 2011. Evolutionary transitions in bacterial symbiosis. Proceedings of the National Academy of Sciences USA. 108: 10800-10807. [PDF]
Sachs, J. L., Skophammer, R.G., and Stajich, J. E. 2014. Evolutionary origins and diversification of proteobacterial mutualists. Proceedings of the Royal Society B. 281:20132146 [PDF]

Microecology of rhizosphere bacteria
Rhizosphere bacteria often encounter multiple plant hosts and diverse environmental pressures in the soil and can exhibit multiple life histories including host-inhabiting and free-living
stages. Research on rhizosphere bacteria -- including pathogens and
beneficial symbionts that inhabit plant roots -- has primarily focused on infection of hosts. In the
mean time, key questions about the ecology and evolution of free-living stages
has remained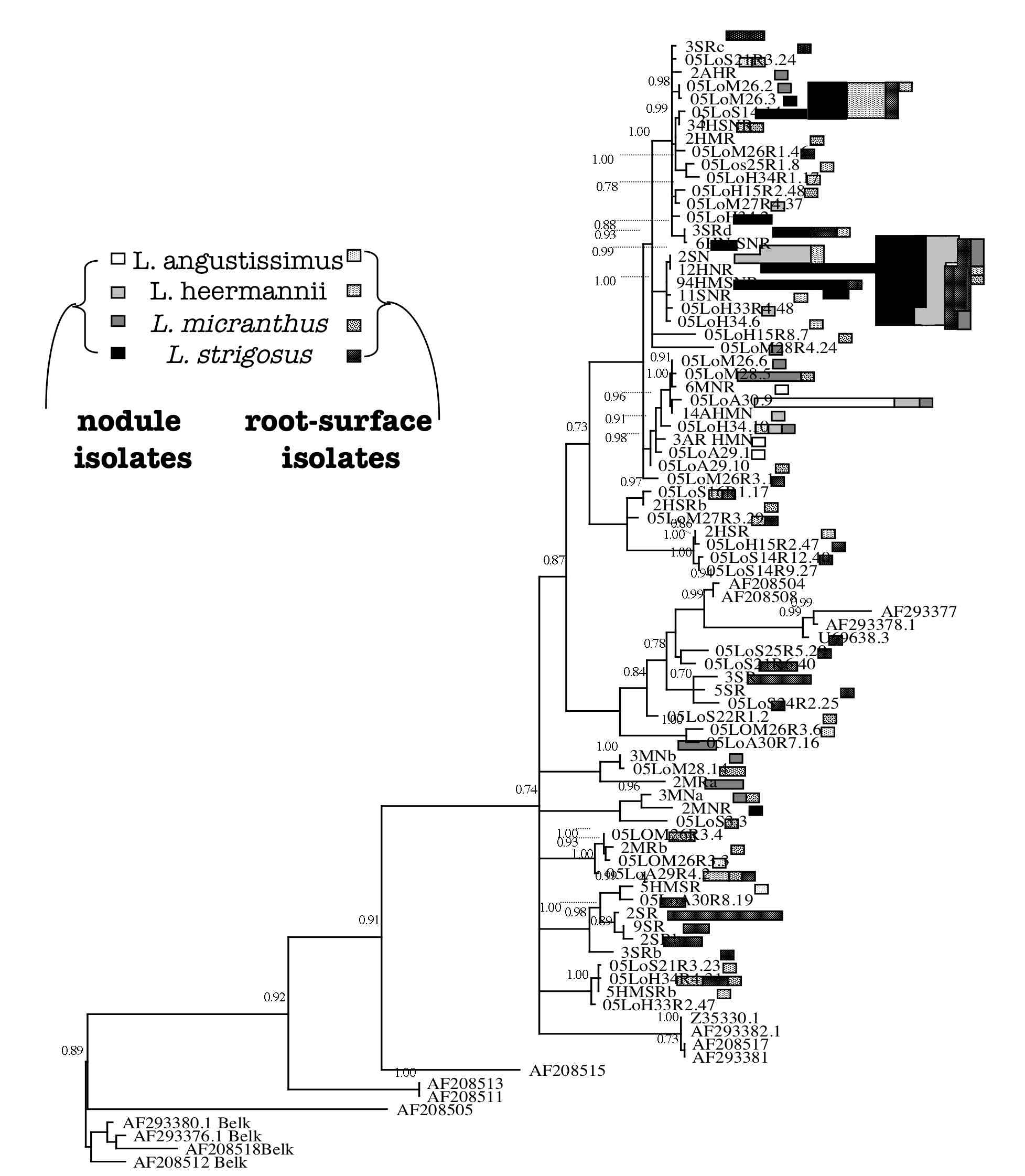  unanswered. For instance, is host association ubiquitous within
bacterial lineages, or do host-infecting genotypes represent subsets of
free-living populations? Assuming that host infection and free-living
existence exert different selective pressures, do diverged bacterial lineages result? Another set of questions addresses the traits that favor the spread of some strains over others in rhizosphere populations. One of the traits that appears to be particularly important is multidrug resistance.
unanswered. For instance, is host association ubiquitous within
bacterial lineages, or do host-infecting genotypes represent subsets of
free-living populations? Assuming that host infection and free-living
existence exert different selective pressures, do diverged bacterial lineages result? Another set of questions addresses the traits that favor the spread of some strains over others in rhizosphere populations. One of the traits that appears to be particularly important is multidrug resistance.
Hollowell, A.C., K.A. Gano, G. Lopez, K. Shahin, J. U. Regus, N. Gleason, S. Greater, V. Pahua, J. L. Sachs. 2015 Native California soils are selective reservoirs for multidrug resistant bacteria Environmental Microbiology Reports. 7,442-449 [PDF]
Sachs, J.L., Kembel, S.W., Lau, A.H., and Simms, E.L. 2009. In situ phylogenetic structure and diversity of wild Bradyrhizobium communities. Applied and Environmental Microbiology 75: 4727-4735. [PDF]